Просмотр исходного кода
Finishes implementing TMVA integration
100 измененных файлов с 896 добавлено и 344 удалено
+ 3
- 0
CMakeLists.txt
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 215
- 0
analysis/MVA_Creation.cpp
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 3
- 3
analysis/MiniTree.hpp
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 37
- 11
analysis/MiniTreeDataSet.hpp
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
analysis/TTTT_Analysis.cpp
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
cmake/FindROOT.cmake
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 162
- 0
docs/MVA__Creation_8cpp.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 16
- 0
docs/MVA__Creation_8cpp__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 0
docs/MVA__Creation_8cpp__incl.md5
|
||
|
||
BIN
docs/MVA__Creation_8cpp__incl.png
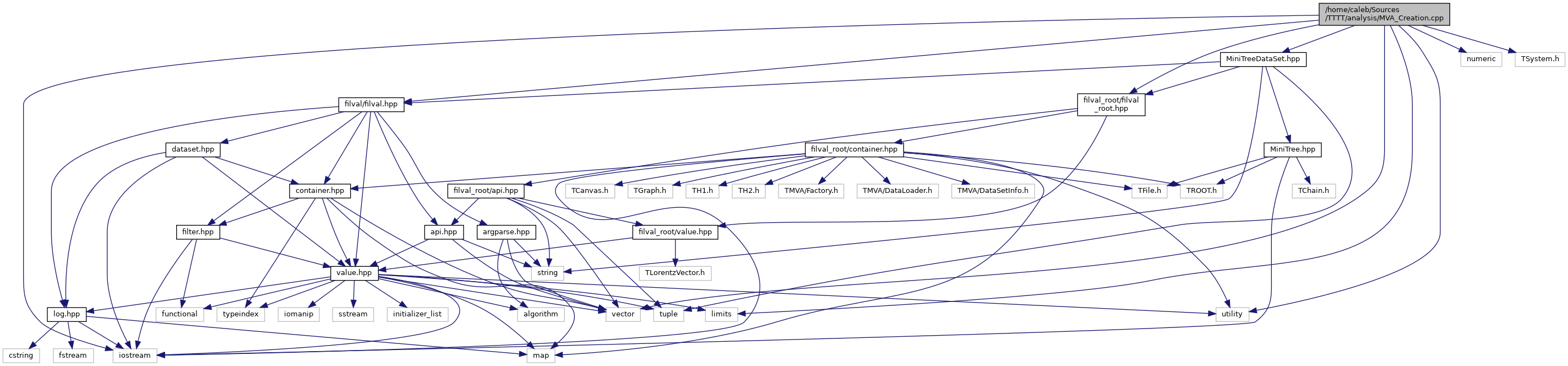
Разница между файлами не показана из-за своего большого размера
+ 82
- 0
docs/MVA__Creation_8cpp_source.html
+ 15
- 14
docs/MiniTreeDataSet_8hpp.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 1
docs/MiniTreeDataSet_8hpp__dep__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/MiniTreeDataSet_8hpp__dep__incl.md5
|
||
|
||
|
||
BIN
docs/MiniTreeDataSet_8hpp__dep__incl.png
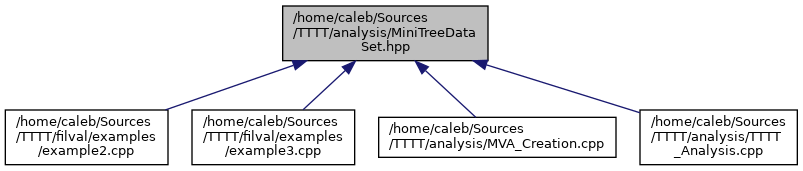
+ 13
- 13
docs/MiniTreeDataSet_8hpp__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/MiniTreeDataSet_8hpp__incl.md5
|
||
|
||
|
||
BIN
docs/MiniTreeDataSet_8hpp__incl.png
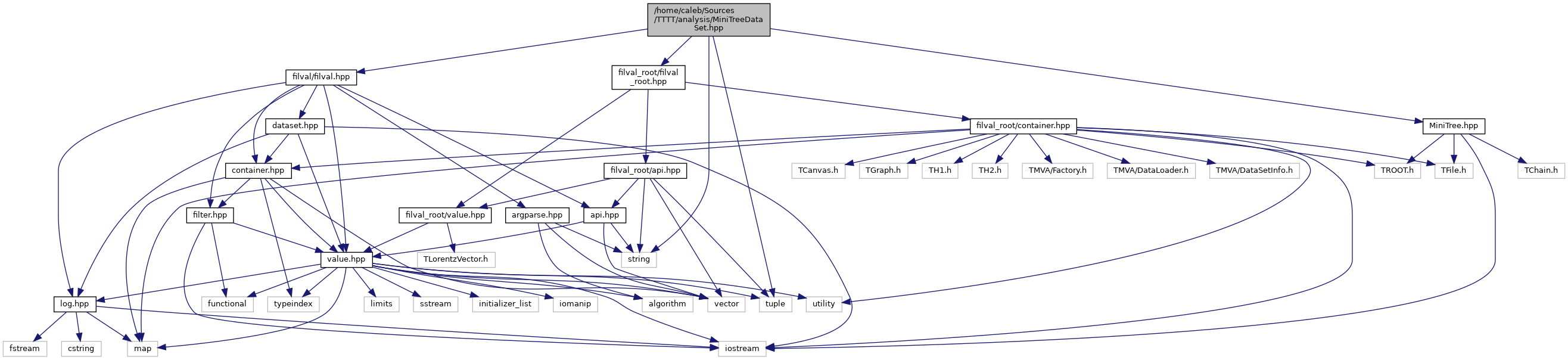
Разница между файлами не показана из-за своего большого размера
+ 2
- 2
docs/MiniTreeDataSet_8hpp_source.html
+ 3
- 2
docs/MiniTree_8hpp.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 3
- 2
docs/MiniTree_8hpp__dep__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/MiniTree_8hpp__dep__incl.md5
|
||
|
||
|
||
BIN
docs/MiniTree_8hpp__dep__incl.png
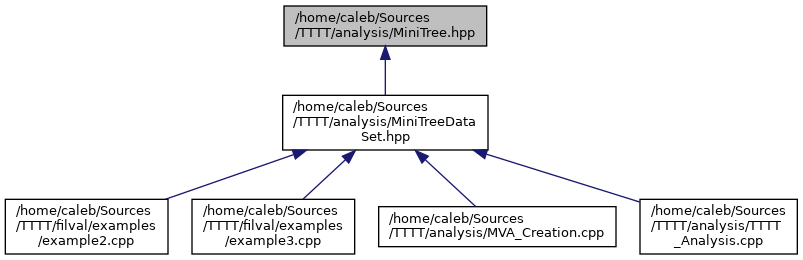
Разница между файлами не показана из-за своего большого размера
+ 1
- 1
docs/MiniTree_8hpp_source.html
+ 2
- 2
docs/TTTT_Analysis.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 15
- 14
docs/TTTT__Analysis_8cpp.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 14
- 14
docs/TTTT__Analysis_8cpp__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/TTTT__Analysis_8cpp__incl.md5
|
||
|
||
|
||
BIN
docs/TTTT__Analysis_8cpp__incl.png
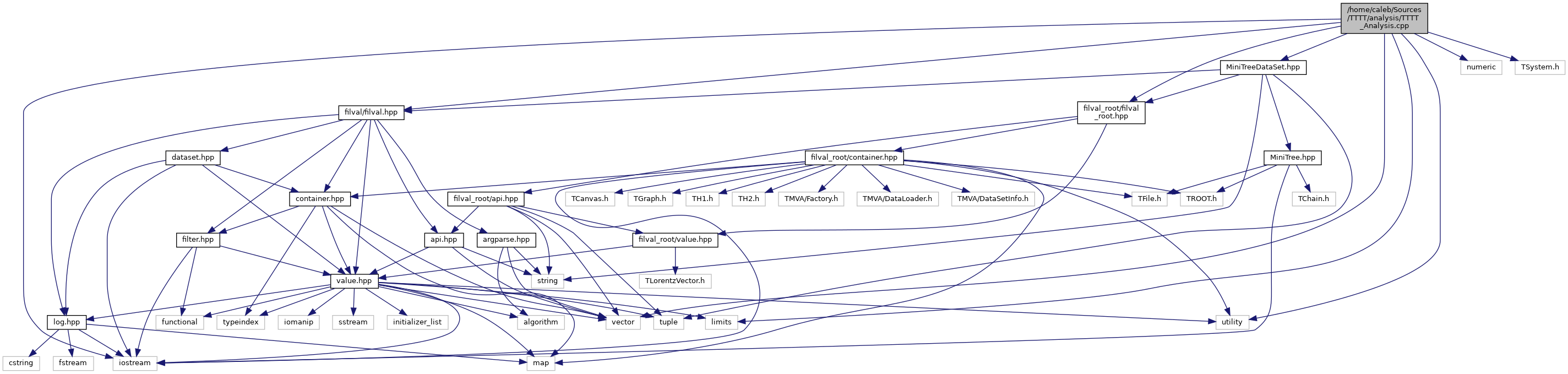
Разница между файлами не показана из-за своего большого размера
+ 1
- 1
docs/TTTT__Analysis_8cpp_source.html
+ 17
- 8
docs/api_8hpp.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 9
- 8
docs/api_8hpp__dep__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/api_8hpp__dep__incl.md5
|
||
|
||
|
||
BIN
docs/api_8hpp__dep__incl.png
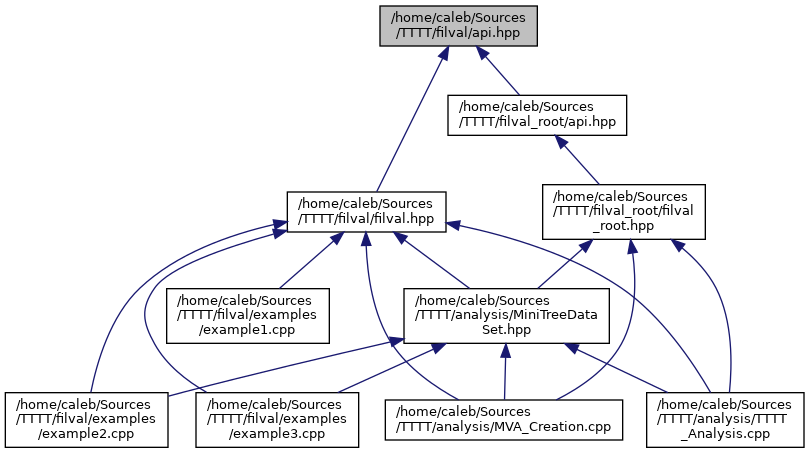
Разница между файлами не показана из-за своего большого размера
+ 17
- 15
docs/api_8hpp_source.html
+ 5
- 4
docs/argparse_8hpp.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 5
- 4
docs/argparse_8hpp__dep__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/argparse_8hpp__dep__incl.md5
|
||
|
||
|
||
BIN
docs/argparse_8hpp__dep__incl.png
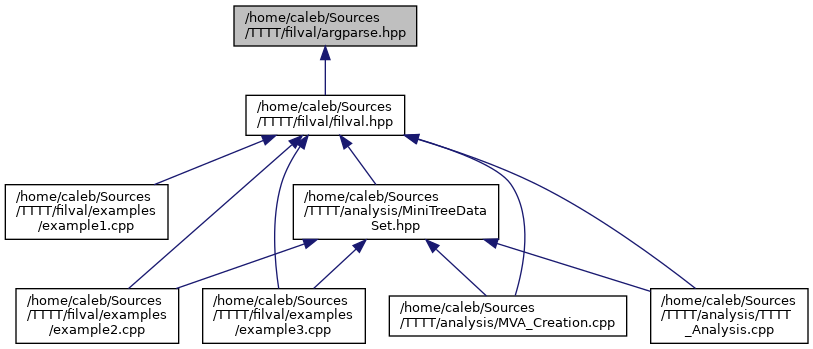
+ 2
- 2
docs/classfv_1_1Apply_3_01Ret_07ArgTypes_8_8_8_08_4.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 15
- 14
docs/classfv_1_1BoundValue-members.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 10
- 3
docs/classfv_1_1BoundValue.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1CartProduct.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1Combinations.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 14
- 13
docs/classfv_1_1ConstantValue-members.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 10
- 3
docs/classfv_1_1ConstantValue.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1Count.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1DeTup.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1DeTupVector.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1ElementOf.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1Filter.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1GenContainer.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1GenContainer__inherit__graph.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1GenContainer__inherit__graph.md5
|
||
|
||
|
||
BIN
docs/classfv_1_1GenContainer__inherit__graph.png
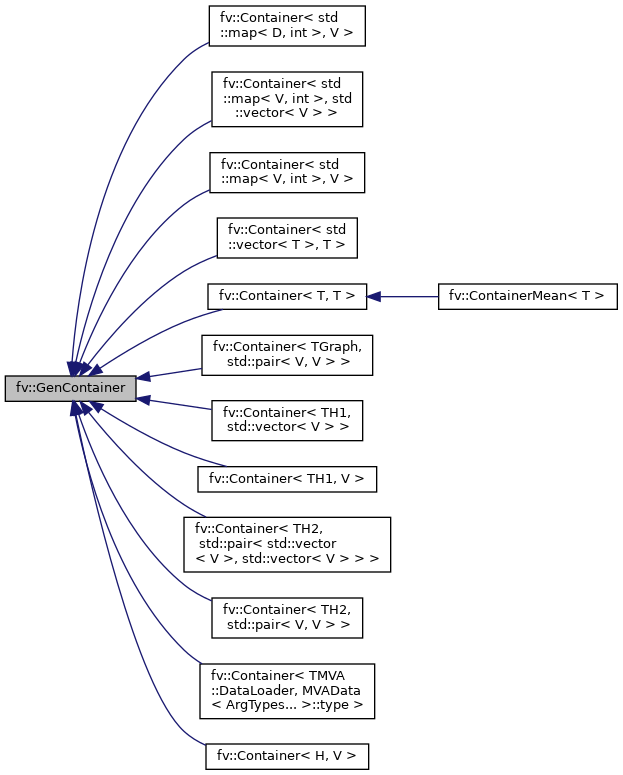
+ 2
- 2
docs/classfv_1_1Map_3_01Ret_07ArgTypes_8_8_8_08_4.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1Max.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1MaxIndex.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1Mean.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1Min.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1MinIndex.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1Pair.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1PointerValue.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1Range.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1Reduce.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1ReduceIndex.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1TupFilter.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 2
- 2
docs/classfv_1_1Tuple.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
Разница между файлами не показана из-за своего большого размера
+ 2
- 2
docs/classfv_1_1WrapperVector.html
Разница между файлами не показана из-за своего большого размера
+ 2
- 2
docs/classfv_1_1Zip.html
+ 1
- 1
docs/classfv_1_1root_1_1Counter.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/classfv_1_1root_1_1CounterMany.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 10
- 9
docs/container_8hpp.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 10
- 9
docs/container_8hpp__dep__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/container_8hpp__dep__incl.md5
|
||
|
||
|
||
BIN
docs/container_8hpp__dep__incl.png
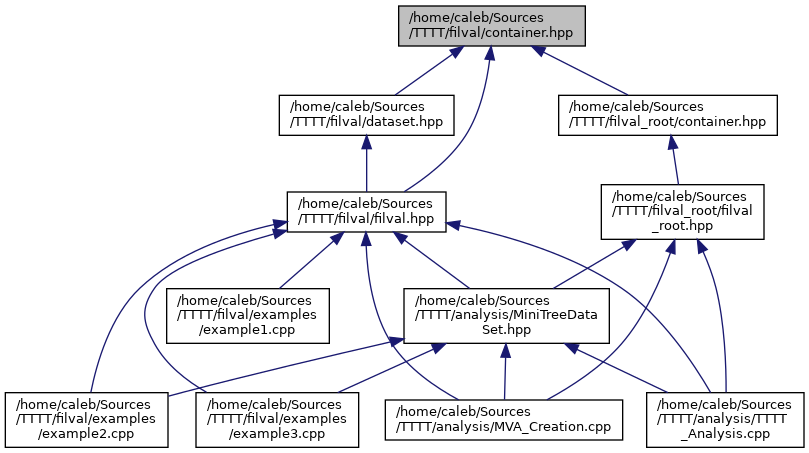
+ 5
- 4
docs/dataset_8hpp.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 5
- 4
docs/dataset_8hpp__dep__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/dataset_8hpp__dep__incl.md5
|
||
|
||
|
||
BIN
docs/dataset_8hpp__dep__incl.png
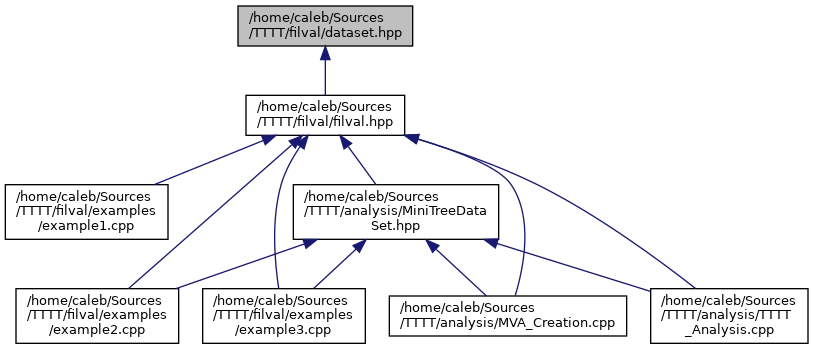
Разница между файлами не показана из-за своего большого размера
+ 1
- 1
docs/dataset_8hpp_source.html
+ 2
- 0
docs/dir_b8678fa8510b7ff9a55ffd4d18fd5e47.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 22
- 21
docs/files.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 11
- 10
docs/filter_8hpp.html
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 11
- 10
docs/filter_8hpp__dep__incl.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/filter_8hpp__dep__incl.md5
|
||
|
||
|
||
BIN
docs/filter_8hpp__dep__incl.png
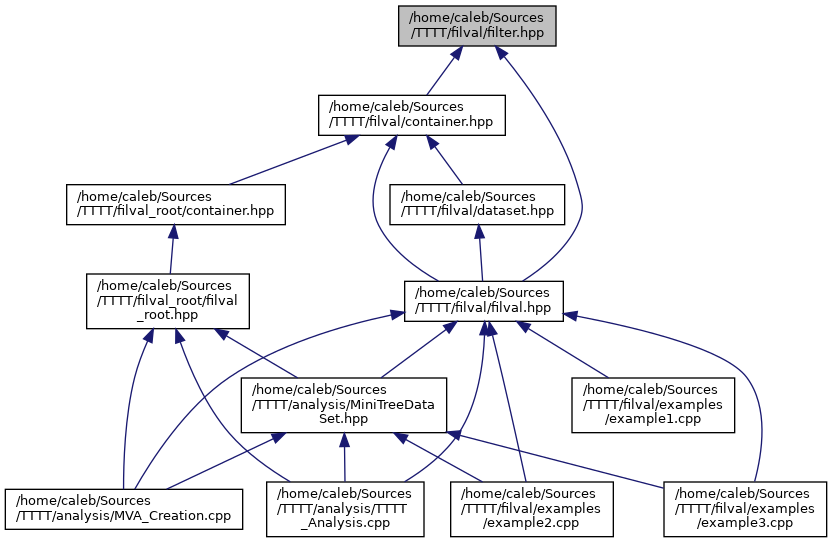
Разница между файлами не показана из-за своего большого размера
+ 58
- 55
docs/hierarchy.html
+ 1
- 1
docs/inherit_graph_0.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/inherit_graph_0.md5
|
||
|
||
|
||
BIN
docs/inherit_graph_0.png
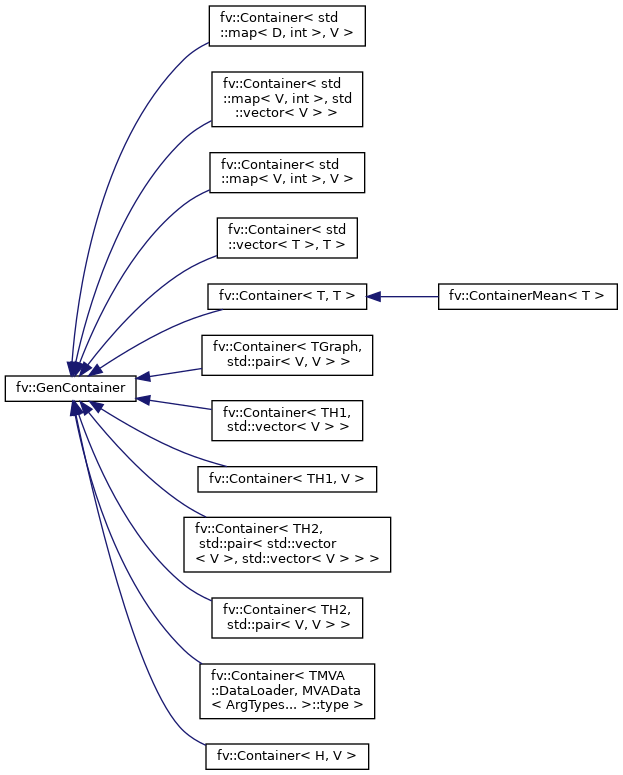
+ 1
- 5
docs/inherit_graph_10.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/inherit_graph_10.md5
|
||
|
||
|
||
BIN
docs/inherit_graph_10.png

+ 5
- 3
docs/inherit_graph_11.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/inherit_graph_11.md5
|
||
|
||
|
||
BIN
docs/inherit_graph_11.png
+ 3
- 1
docs/inherit_graph_12.map
|
||
|
||
|
||
|
||
|
||
|
||
|
||
+ 1
- 1
docs/inherit_graph_12.md5
|
||
|
||
|
||
+ 0
- 0
docs/inherit_graph_12.png
